Contextual data models in health AI integrate information from wearables, lab results, and lifestyle data to provide a complete view of patient health. This approach addresses fragmented systems and improves diagnostic accuracy by combining data sources like electronic health records (EHRs), IoT devices, and genetic profiles. For example, blending heart rate data with sleep patterns or lab results can reveal insights that single-source analysis misses.
Key points:
- Data sources: Wearables (e.g., heart rate, sleep), lab results (e.g., imaging, biomarkers), and lifestyle data (e.g., diet, activity).
- Fusion techniques: Sensor-level (raw data), feature-level (extracted metrics), and decision-level (model outputs).
- Challenges: Data fragmentation, bias, and interoperability issues.
- Standards: FHIR and OMOP frameworks streamline data sharing while maintaining privacy and security.
- Privacy tools: Federated learning and differential privacy protect sensitive information during data integration.
553 - How To Turn Healthcare Data into AI-Driven Action
Key Data Sources for Contextual Fusion
Health AI draws on three main data streams to create a well-rounded view of your health. Each source adds its own layer of insight, and when combined, they enable AI systems to go beyond one-size-fits-all advice, offering tailored recommendations instead. Let’s break down these essential data streams and how they work together to provide a full picture of patient health.
Wearables and Biosensors
Devices like smartwatches, fitness trackers, and other wearables generate real-time physiological data that traditional healthcare often misses. These gadgets continuously monitor critical metrics such as HRV, stress, and sleep tracking, blood oxygen levels (SpO2), and even electrocardiogram (ECG) readings. Some wearables go a step further - smart insoles track gait and balance, while respiratory bands measure breathing patterns during activities or rest [5].
For instance, a study involving over 139 million heart rate measurements from smartwatches helped train AI models to detect atrial fibrillation [8]. This kind of high-resolution data enables AI to pick up on subtle changes, like early signs of stress or inflammation, long before symptoms appear. However, to ensure accuracy, these devices need regular calibration to avoid "data drift", which can happen due to environmental factors or device wear over time [5].
When wearable data is combined with clinical and behavioral information, it becomes far more powerful, offering a richer context for understanding health trends.
Lab Results and Diagnostic Data
Clinical lab tests and medical imaging provide the biological foundation needed to make sense of real-time sensor data. This includes structured information from blood tests, biomarker panels, genomic profiles, and imaging data like MRI, PET, CT scans, and X-rays [6][7]. These insights help AI distinguish between normal variations and potential health issues.
For example, a chest X-ray showing lung opacity could indicate anything from pneumonia to lung cancer. When paired with lab results and patient history, the diagnosis becomes much clearer [6]. A survey revealed that 87% of radiologists believe clinical context significantly improves their image interpretations [6].
The benefits of integrating multiple data types are clear. In 2019, researchers developed a breast cancer risk model that combined mammograms with non-imaging data like age, family history, and BRCA mutation status. This approach achieved an AUROC of 0.70, outperforming models that relied solely on mammograms (0.68) or risk scores (0.67) [6]. Similarly, a 2018 study used MRI data alongside clinical and genetic information to predict Alzheimer’s disease progression, achieving a perfect AUROC of 1.00 [6]. These examples highlight how blending imaging, lab results, and other data enhances predictive accuracy.
When this clinical data is integrated with real-time sensor readings and behavioral patterns, it forms a robust foundation for personalized health strategies.
Lifestyle and Behavioral Data
Patient-generated health data (PGHD) fills in the gaps by explaining the why behind physiological changes. This includes information about diet, exercise habits, medication adherence, mental health, social media activity, location, and environmental exposures [7]. For example, a spike in heart rate might make sense if it coincides with a recorded workout session.
In one 2018 study, researchers analyzed metabolome, microbiome, genetic, and imaging data using unsupervised learning. They identified biomarker patterns that detected diabetes more accurately than traditional metrics like glucose levels or insulin resistance [7]. This demonstrates how lifestyle data can uncover trends that might otherwise go unnoticed.
When these three data streams come together, they enable what experts call "deep phenotyping" - a method of categorizing patients based on the interplay between genetics, environment, and behavior [7]. Contextual fusion turns this mountain of data into actionable insights by connecting the dots across different sources.
| Data Source | Key Metrics | Primary Contribution |
|---|---|---|
| Wearables | HRV, SpO2, sleep patterns, gait metrics | Real-time health trends |
| Lab Results | Biomarkers, genomics, imaging data | Biological foundation and clinical depth |
| Lifestyle Data | Diet, activity, medication adherence | Behavioral context and intervention tracking |
Techniques for Contextual Data Fusion
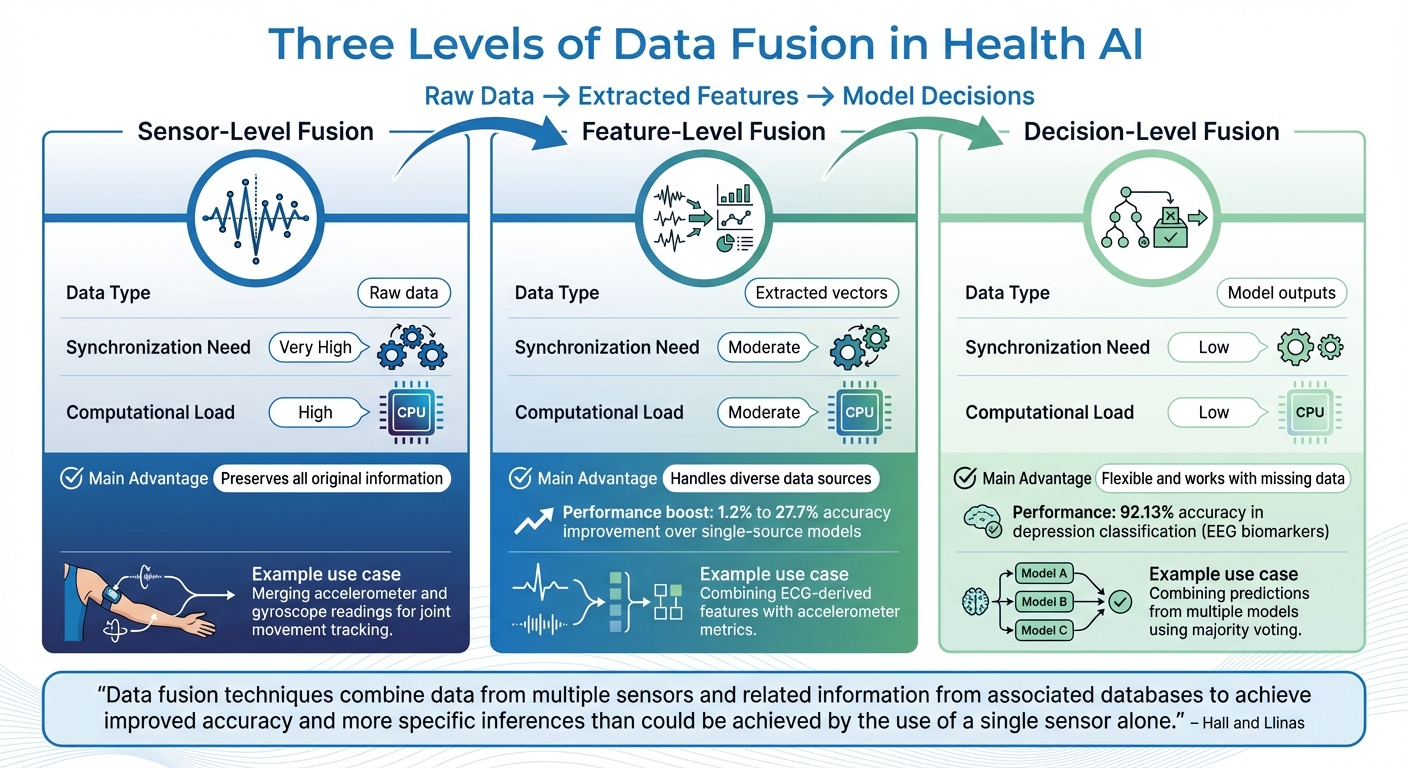
Three Levels of Data Fusion in Health AI: Comparison of Sensor, Feature, and Decision-Level Techniques
By combining data from wearables, lab results, and lifestyle inputs, contextual data fusion creates rich, in-depth analyses. The method you choose depends on your goals and the type of data you're working with. Health AI systems often rely on three main approaches to integrate information, each with its own strengths and limitations.
Sensor-Level, Feature-Level, and Decision-Level Fusion
Sensor-level fusion (or signal-level fusion) operates directly on raw data from sensors. This method works best when combining data from similar sensors, like pairing accelerometer and gyroscope readings to track joint movements with precision[9]. However, it demands significant computational resources and is only practical when sensors capture the same type of signals. For example, merging raw ECG data with text-based notes isn’t feasible using this approach[5].
Feature-level fusion takes another path. Instead of raw data, it focuses on extracting key characteristics - like average heart rate, step count variability, or features derived from neural networks - and merges these into a unified feature set[9][14]. This method is ideal for integrating diverse data types, such as ECG-derived features and accelerometer metrics. Studies show that multimodal fusion models using this technique can improve accuracy by 1.2% to 27.7% compared to single-source models[14]. These feature sets then guide the selection of machine learning models, which refine the data into actionable clinical insights.
Decision-level fusion (also called late fusion) comes at the end of the pipeline. Here, individual models analyze data from each source separately and produce their own predictions. These outputs are then combined using methods like majority voting or Bayesian inference[11][13]. This approach is particularly useful for asynchronous data or when data availability varies, as it remains effective even with intermittent inputs[13][14]. For example, researchers used decision-level fusion to classify depression by combining linear and non-linear EEG biomarkers, achieving an impressive 92.13% accuracy[10].
"Data fusion techniques combine data from multiple sensors and related information from associated databases to achieve improved accuracy and more specific inferences than could be achieved by the use of a single sensor alone." – Hall and Llinas [13]
A more advanced technique, joint fusion, leverages deep learning by merging features from intermediate layers of neural networks. Unlike early fusion methods, joint fusion allows for error refinement through backpropagation, enhancing feature extraction over time. This approach has shown exceptional results, such as achieving a perfect AUROC score in predicting Alzheimer's progression[14].
| Fusion Level | Data Type | Synchronization Need | Computational Load | Main Advantage |
|---|---|---|---|---|
| Sensor (Signal) | Raw data | Very High | High | Preserves all original information [9][10] |
| Feature | Extracted vectors | Moderate | Moderate | Handles diverse data sources [9][14] |
| Decision | Model outputs | Low | Low | Flexible and works with missing data [13][14] |
Machine Learning Models for Data Fusion
Once the data is fused, the next step is selecting the right machine learning models to turn it into actionable insights. Convolutional Neural Networks (CNNs) are excellent for extracting features from medical images like MRIs or CT scans, while Recurrent Neural Networks (RNNs) and LSTMs excel at processing time-series data from wearables or sequential clinical notes[10][14]. For instance, researchers at Northumbria University used CNNs to analyze multi-location accelerometer and ECG data, identifying arrhythmias by first determining whether the individual was sitting or walking[5].
For structured data, Random Forests and Support Vector Machines (SVMs) are solid choices. Random Forests not only perform well in classification tasks, but they also offer interpretability, helping clinicians understand the reasoning behind a model's predictions[10][12]. Similarly, SVMs are effective in high-dimensional scenarios, such as distinguishing between malignant and benign tumors using fused biomarker panels[12][13].
For real-time applications, Kalman Filters and Particle Filters are often used in sensor-level fusion, particularly in motion tracking and biomechanics[12]. Meanwhile, Bayesian Inference models shine in decision-level fusion by handling uncertainty effectively. They combine prior probabilities with new evidence to produce calibrated predictions. In a COVID-19 screening study, researchers used a multimodal framework that integrated lab tests, clinical records, and CT scans, demonstrating how Bayesian methods can handle diverse data types under uncertain conditions[10].
"Selecting the appropriate techniques based on the desired outcome is crucial for successful implementation." – Yunus Celik, Northumbria University [9]
Different models serve different purposes: CNNs for imaging, LSTMs for sequential data, Random Forests for structured records, and Bayesian models for uncertainty management. Together, they transform fused datasets into insights that can enhance patient care.
Interoperability and Standards in Health AI
Interoperability in health AI helps break down silos, streamlining data sharing and cutting down on administrative inefficiencies. By harmonizing data across systems, it reduces delays and ensures that critical information is accessible when needed. Without standardized practices, up to 30% of collected data could be lost, and projects might face delays of up to six months[3][15]. To make context-aware data fusion effective, systems need to "speak the same language."
Health Data Standards: FHIR and OMOP

FHIR (Fast Healthcare Interoperability Resources) is a flexible, resource-based framework for exchanging health data. Instead of transferring entire documents, FHIR organizes information into smaller, modular resources - like Patient, Observation, or Medication - that can be accessed in real-time through RESTful APIs[15]. This modular design is perfect for AI systems requiring specific data points on demand. For instance, an AI might pull recent lab results to adjust patient-centered treatment plans or retrieve heart rate variability trends to refine sleep recommendations.
"The move to Fast Healthcare Interoperability Resources (FHIR) isn't just about adopting new technology; it's about unlocking a new era of efficiency and revenue." – Sony George, blueBriX[15]
OMOP (Observational Medical Outcomes Partnership), on the other hand, focuses on standardizing data into relational tables tailored for research and large-scale analysis. While FHIR is ideal for transactional, real-time data needs, OMOP excels in batch processing and creating datasets ready for analytics. Together, these frameworks complement each other: FHIR supports clinical decision-making and mobile health apps, while OMOP is better suited for training AI models using large observational datasets.
To ensure smooth data integration across systems, semantic normalization is a must. This involves mapping local codes to universal standards like SNOMED CT for conditions, LOINC for lab tests, and RxNorm for medications[15][3]. Without this step, inconsistencies can arise - for example, one system might label "systolic BP", while another uses "SBP", leading to mismatched insights. By focusing on the clinical meaning rather than just the format, concept-based harmonization ensures that mappings are reusable and scalable across platforms[3].
| Standard | Primary Use Case | Data Structure | Exchange Method |
|---|---|---|---|
| FHIR | Real-time data exchange, EHR integration | Modular resources (JSON/XML) | RESTful APIs |
| OMOP | Observational research, population analysis | Standardized relational tables | Batch/Analytics-ready |
| C-CDA | Care transitions, legal documentation | Monolithic XML documents | File-based (SFTP, secure email) |
While these standards enable seamless data flow, they also bring privacy concerns to the forefront.
Privacy and Ethical Considerations
Using FHIR and OMOP for standardized data exchange can improve efficiency, but it also highlights the need to address privacy concerns. When data moves between wearables, labs, and AI systems, protecting sensitive information becomes crucial. Technologies like OAuth 2.0 and OpenID Connect play a key role in securing these exchanges, ensuring that only authorized systems can access patient data[15][8].
Federated learning is a promising approach to enhance privacy. Instead of centralizing raw patient data, this method trains AI models locally on devices or within institutions, then combines only the model updates. Studies show that secure, multi-modal federated learning can improve diagnostic accuracy by 9.8% and reduce client dropout rates by over 50%, all while safeguarding privacy[1].
Differential privacy adds another safeguard by introducing controlled noise into datasets. This prevents the reconstruction of individual patient records while maintaining accuracy for clinical purposes[1]. Implementing granular access controls ensures that AI agents only access the data they need. For example, a sleep optimization tool might require heart rate variability and activity logs but wouldn’t need access to lab results or prescription histories.
A patient-first approach to consent is critical. Individuals should have control over which AI systems access their data and how it’s used[7][2]. Building transparency into data usage fosters trust and ensures that AI systems are deployed ethically and responsibly.
sbb-itb-f5765c6
How BondMCP Enables Context-Aware Health Optimization
Fragmented health data has long been a hurdle in achieving seamless health optimization. Wearables monitor sleep, labs analyze biomarkers, and fitness apps track workouts - but none of these systems talk to each other. BondMCP tackles this issue by acting as a universal interface that connects these disparate systems[2]. Instead of complex, one-off integrations for every connection, BondMCP simplifies the process, reducing the integration complexity from M×N to M+N. This streamlined approach enables ecosystem-wide interoperability[2].
Shared Context Layer and Health Ontology
At the core of BondMCP is its shared context layer, which brings together clinical imaging, electronic medical records (EMRs), and real-time data from wearables into a unified framework[1]. This system organizes data into three functional categories:
- Resources: Read-only data like lab results or wearable logs.
- Tools: Functions that can perform tasks, such as querying a database or sending alerts.
- Prompts: Predefined templates that guide AI through specific clinical workflows.
By standardizing data access and interpretation, BondMCP ensures compatibility across devices. For instance, a heart rate reading from an Apple Watch is seamlessly aligned with data from a Fitbit, eliminating the inconsistencies that often arise in traditional systems.
"Models are only as good as the context we provide them." – Charles Shen, PhD, EMBA[4]
BondMCP uses JSON-RPC 2.0 for communication, supporting both local (stdio) and remote (HTTP + Server-Sent Events) transports[2]. This setup gives developers the flexibility to either keep sensitive data on-device for privacy or stream real-time updates from cloud-based sources. The result? A unified framework that simplifies development while enabling secure and adaptable health applications.
Plug-and-Play SDK for Developers
BondMCP's SDK builds on the shared context layer to make integration straightforward for developers. Instead of reworking core systems for every new use case, developers can use high-level tools like FastMCP and decorators such as @mcp.tool() to auto-generate input schemas and descriptions directly from function docstrings[2]. This means a developer can quickly wrap an existing database or wearable API into an MCP server with minimal effort, making it accessible to any MCP-compliant AI system.
The SDK also supports dynamic capability discovery, which allows AI agents to automatically identify available tools and data sources without requiring manual updates[16][17]. Security is a priority here, with features like OAuth 2.0/2.1 authentication, end-to-end encryption, and granular access controls ensuring HIPAA compliance[16][17][18]. For example, a sleep optimization agent might access heart rate variability and activity logs but would be restricted from viewing prescription histories. This built-in security ensures sensitive health data remains protected while enabling advanced AI-driven insights.
Scalable Solutions for Clinics and Biohackers
BondMCP extends its benefits to both clinical settings and personal health enthusiasts. For clinics, it enables federated learning, allowing multiple institutions to train AI models collaboratively without centralizing patient data[1]. This approach meets stringent HIPAA and GDPR requirements while leveraging large datasets for research. Additionally, its energy-efficient scheduling reduces client dropout rates by 54%, ensuring stable participation in remote or resource-limited environments[1].
For biohackers, BondMCP eliminates the hassle of manually syncing data across various apps. AI agents can automatically pull together lab results, wearable metrics, and supplement logs into a single, actionable dashboard. These agents can go beyond simply displaying data - they can perform tasks like calculating personalized health scores or updating records using standardized function calls[2]. This transforms static data into a dynamic, real-time health management system that adapts to new information as it becomes available.
Challenges and Best Practices for Contextual Data Fusion
Combining health data is no easy task. In fact, as much as 30% of data might be discarded simply because it can't be reconciled. The hurdles are numerous: incompatible formats like CSV, JSON, and XML, or comparing agent protocols like MCP and A2A; semantic ambiguity (for example, does "blood pressure" mean systolic, diastolic, or an average?); and varying coding systems such as ICD, SNOMED CT, and LOINC. Context also matters - a diagnosis made in an emergency room might carry a completely different weight than one recorded during a controlled clinical trial. Losing this context can lead to flawed AI outputs, which is a serious concern [3].
"Attempting to combine these datasets without proper harmonization leads to errors, bias, and wasted resources." – Maria Chatzou Dunford, CEO and Co-founder, Lifebit [3]
The solution begins with ensuring strong data hygiene.
Data Hygiene and Validation
High-quality data is the foundation for reliable AI outcomes. Unfortunately, issues like missing values, inconsistent granularity (e.g., "one pill" versus "500 mg"), typos in patient identifiers, and mismatched date formats can quickly derail any harmonization effort. That’s why data profiling is a critical first step. It helps identify value distributions, completeness, and potential gaps in the dataset.
A concept-based approach can also help. Instead of focusing on specific formats (e.g., "MM/DD/YYYY" for a date), this method emphasizes the underlying clinical meaning - like treating "date of birth" as the core concept. Tools like SSSOM (Simple Standard for Sharing Ontological Mappings) can make mappings more transparent and easier to audit by adding metadata such as confidence levels and justifications for the mappings. On-the-fly mappings, which preserve both data integrity and provenance, further enhance the process. Together, these tactics resolve ambiguities and maintain clinical accuracy [3].
When done right, these practices ensure that the merged data accurately reflects the original clinical context.
Bias Mitigation in Fused Data
Bias is another significant challenge in data fusion, especially when certain populations are underrepresented or excluded altogether. A well-documented example is a skin cancer detection algorithm that performed poorly for darker-skinned individuals because it was trained primarily on data from light-skinned patients [3].
To address this, datasets need to be harmonized across diverse demographics and geographic areas. Tracking variables like gender, ethnicity, and socioeconomic status can help identify and correct for historical recruitment biases [7]. Federated learning offers another solution by training AI models on decentralized data, ensuring global representation while maintaining privacy [1][3]. For systems involving wearables, energy-aware scheduling can minimize "client dropout" among users with older devices or limited internet bandwidth. Research shows that this approach can cut dropout rates by 54% and improve diagnostic accuracy by 9.8% compared to standard federated learning models [1].
Conclusion: Building the Future of Context-Aware Health AI
The future of health AI hinges on breaking down data silos and building systems capable of seeing the entire clinical picture. With healthcare data exploding from 153 exabytes in 2013 to a staggering 2,314 exabytes in 2020, the need for smarter data integration strategies has become impossible to ignore [19]. Gathering data alone isn’t enough; the real game-changer lies in combining inputs from wearables, lab tests, lifestyle habits, and clinical records into a cohesive, actionable framework.
"The real breakthrough in health data harmonization is treating it as a translation problem, not a formatting problem. The most powerful mappings preserve what the data actually means." – Maria Chatzou Dunford, CEO and Co-founder, Lifebit [3]
This challenge of unifying diverse data streams has opened the door for solutions like BondMCP.
By focusing on seamless interoperability and a shared context layer, BondMCP tackles this challenge head-on. It uses a health-specific ontology that enables AI systems to communicate effortlessly. Instead of creating custom connectors for every data source - a process that often spirals into chaos - BondMCP simplifies the equation. Developers only need to implement the protocol once, unlocking access to an entire ecosystem [2]. On a practical level, this means your sleep tracker, fitness coach, and supplement regimen can finally work in harmony. For clinics and biohackers, it offers a scalable approach to precision health without relying on clunky dashboards or isolated tools.
When organizations focus on data hygiene, semantic alignment, and bias reduction, they turn AI from a theoretical concept into a powerful tool for health optimization. Those who invest in proper data preparation avoid the pitfalls of poorly harmonized systems - like six-month delays or losing 30% of their data to inefficiencies [3]. Done right, multi-modal data fusion can boost diagnostic accuracy by up to 9.8% and cut device dropout rates by 54% through smarter energy management [1]. These aren’t just incremental improvements - they mark the line between AI that merely exists and AI that transforms health outcomes on a large scale.
FAQs
How does combining different types of health data improve diagnostic accuracy in AI systems?
Combining various health data sources - like medical records, imaging, genomics, and wearable sensor data - into a single framework allows AI systems to pick up on patterns that might go unnoticed when analyzing each source separately. This approach, called contextual data fusion, organizes and aligns these inputs to minimize noise, clarify ambiguities, and provide more accurate diagnoses. For example, integrating imaging data with lab results and medication history enables AI to cross-check findings, which can enhance accuracy and reduce false positives for conditions such as cancer or heart disease.
This technique also plays a key role in tailoring care to individual patients by offering a complete picture of their health. By combining data on activity levels, sleep patterns, and lab results with diagnostic imaging, AI can differentiate between disease-related changes and normal fluctuations. Tools like the BondMCP Health Model Context Protocol make this process smoother by standardizing how data is interpreted and allowing real-time synchronization. The result? Clinicians get sharper, more actionable insights to guide patient care.
What are the biggest challenges in combining health data from different sources?
Integrating health data from various sources - like wearables, electronic health records (EHRs), lab results, and patient-generated data - presents a host of challenges. One of the biggest hurdles is the incompatibility of data formats and standards. Wearables, for instance, often rely on proprietary data models, while clinical systems adhere to standards like HL7 FHIR. Bridging these differences requires extensive data mapping, a process that’s not only time-consuming but also prone to potential data loss.
Another significant issue is the lack of interoperability. Many consumer health apps and devices operate in silos, making it difficult to integrate patient-generated health data (PGHD) with clinical systems. This fragmentation forces healthcare providers to navigate multiple tools, increasing the likelihood of overlooking critical insights.
Lastly, there’s the challenge of merging multimodal data streams. Combining data like heart rate, sleep patterns, and activity levels demands advanced techniques to ensure consistency and accuracy, which is no small feat.
The BondMCP – Health Model Context Protocol tackles these challenges head-on by offering a shared context layer. This layer harmonizes data, ensures interoperability, and enables real-time AI-driven insights, creating a unified, privacy-conscious system tailored for personalized health management and clinical application.
How does federated learning protect patient privacy in health AI systems?
Federated learning safeguards patient privacy by ensuring that raw health data stays where it originates - on devices like wearables or within systems such as electronic health records (EHRs). Instead of sending sensitive information to a central server, each system trains its own local AI model using its own data. Only encrypted model updates are shared with a central aggregator, reducing the risk of exposing personal health details and complying with privacy regulations like HIPAA in the U.S.
To make this process even more secure, federated learning incorporates advanced methods like secure multiparty computation, homomorphic encryption, and differential privacy. These techniques ensure that even the shared model updates cannot be reverse-engineered to extract individual data. This approach allows AI systems in healthcare to train on diverse, high-quality datasets - like data from wearables, lab tests, and medical imaging - while maintaining strict confidentiality.
By keeping data decentralized and prioritizing privacy, federated learning builds trust in AI-powered healthcare and supports the creation of more personalized and context-aware models.